BIOINFORMATIC SERVICES
Welcome to our bioinformatics services! We are a leading provider of cutting-edge bioinformatics solutions tailored to meet the diverse needs of researchers, scientists, and organizations in the life sciences. With our deep expertise in computational biology, data analysis, and genomics, we are dedicated to helping you extract meaningful insights from complex biological data. Whether you require genome analysis, metagenomics, transcriptomics, proteomics, or other bioinformatics services, our team of skilled bioinformaticians and data scientists is here to support your research goals. From data processing and analysis to interpretation and visualization, we offer comprehensive and reliable solutions to drive scientific discoveries, accelerate research, and advance precision medicine. Partner with us and unlock the full potential of your biological data with our cutting-edge bioinformatics services.If you do not see what you are looking for below, please reach out to us at bioinformatics.core@tamu.edu to set up a free initial consultation:
Metagenomics
Description: We offer analysis of metabarcoding data (16S, ITS, and custom amplicons) as well as assembly, annotation, and analysis of shotgun metagenomic data from short- and long-read DNA sequencing data. In addition, we offer 5-week, 1-credit workshops on analysis of shotgun metagenomic and metabarcoding data each fall and spring semester, respectively.
Tools: qiiime2, MetaPhlAn, Kraken, MetaSpades, Megahit, MetaBAT, and BV-BRC pipelinesTools: DIAMOND, Kraken, MetaSpades, MetaBAT, (get list from Chris)
Genome Assembly & Annotation
Description: We offer de novo assembly and annotation of animal, plant, bacterial, and viral genomes from short-read , long-read and hybrid data sets using gold-standard open source tools. We also perform optical genome mapping using the Bionano Saphyr system for optical map guided genome assembly with or without Hi-C data.
Tools: Canu, Flye, MaSuRCA, SPades, Megahit, BUSCO, QUAST.

Comparative Genomics
Description: With state-of-the-art open source tools and methodologies, we analyze genomic data from multiple sources, including whole-genome sequencing, transcriptomics, and functional genomics data. Our bioinformatics pipelines are designed to process and integrate diverse genomic datasets, enabling us to identify orthologous genes, predict gene functions, analyze genomic rearrangements, detect genetic variations, and assess the evolutionary relationships between genomes.
Tools: OrthoFinder, IQTREE, EDTA, CACTUS, Circos, ANI/AAI, Phylogenomics
Single Cell RNA-Seq Analysis (scRNA-Seq)
Description: We preprocess raw scRNA-seq data, perform quality control assessments, normalize expression values, and identify differentially expressed genes. Our pipelines incorporate algorithms for dimensionality reduction, clustering, cell type identification, trajectory analysis, and visualization, enabling comprehensive exploration of cellular heterogeneity and differentiation processes. We offer a range of services tailored to meet your specific research needs. Whether you require data preprocessing and quality control, cell type identification and annotation, differential gene expression analysis, trajectory inference, or integration of multiple datasets.
Tools: Cell Ranger, Space Ranger, Seurat, PCA, tSNE
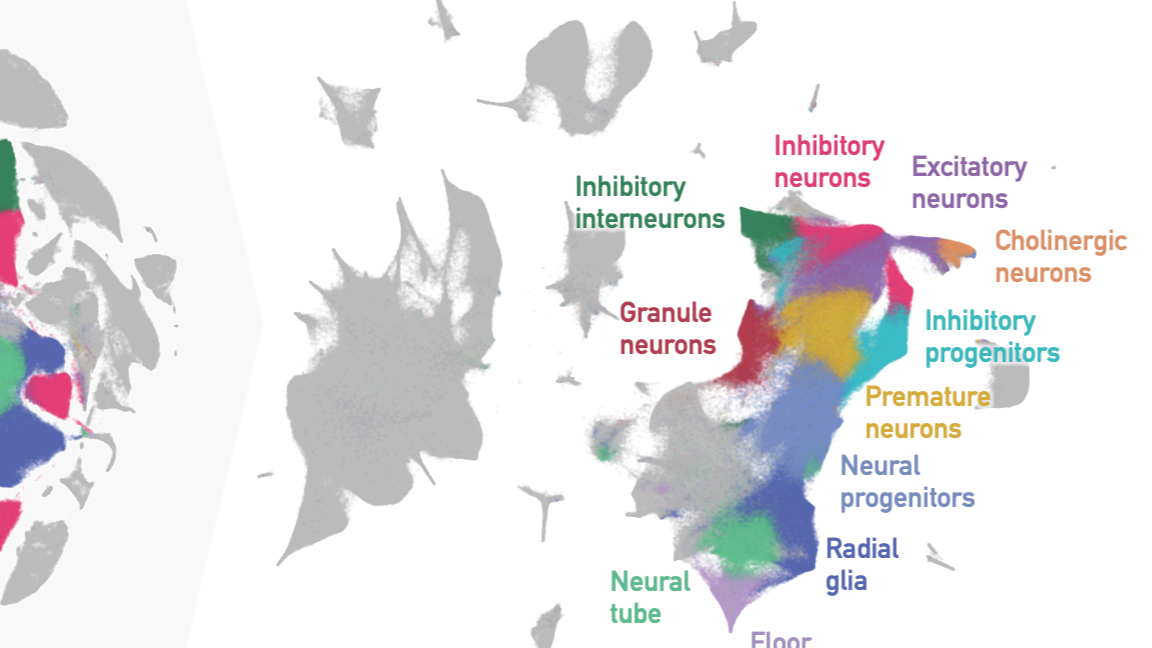
Total & mRNA-Seq Analysis
Description: Whether it’s gene expression patterns, identifying differentially expressed genes, exploring alternative splicing events, or unraveling complex regulatory networks, our comprehensive RNA-seq analysis services provide you with the expertise and tools needed to extract meaningful and actionable information from your transcriptomic data. Partner with us to unlock the full potential of your RNA-seq data and gain a deeper understanding of the molecular mechanisms that underlie biological processes.
Tools: Salmon, kallisto, DESeq2, EdgeR, HISAT2, BioPython
Custom Pipelining & Analysis
Description: We understand that every research project has unique requirements and demands tailored analysis approaches. That’s why we offer custom pipelining solutions, designed to meet your specific bioinformatics needs. Whether you need data preprocessing, quality control, statistical analysis, machine learning, or visualization, our custom pipelines are designed to streamline and automate your analysis workflow, saving you time and effort.
Languages: R, Python, Bash, Perl

